Alternative nucleic acid amplification methods: part 1
Editor’s note: Back by popular demand, here is the first installment of the second volume of “The Primer: A Guide to Molecular Diagnostics.” Same as in 2013, John Brunstein, PhD, presides over this progressive series of columns—a kind of mini-textbook covering important topics in MDx.
Over the course of most of last year’s run of The Primer, our coverage was primarily on PCR- and RT-PCR-based amplification methods. While that focus is reflective of the frequent application of these methods in the clinical laboratory setting, PCR is not the only method of nucleic acid amplification and detection that is in use today. Other approaches, each with its own strengths and weaknesses, are also currently employed in some settings, and they will likely continue to be methods of choice in some applications. Over the course of the next two installments of The Primer, we’ll direct our attention to some of these that laboratorians are likely to encounter—now or in the future.
The acronym “NASBA” stands for nucleic acid sequence-based amplification, and that covers a few variations on techniques using RNA transcription as part of the amplification cycle. Perhaps best known and most common among NASBA methods is transcription mediated amplification or TMA, developed originally in the mid-1990s as a methodology for detection of RNA viruses. In essence, the approach works as outlined in Figure 1.
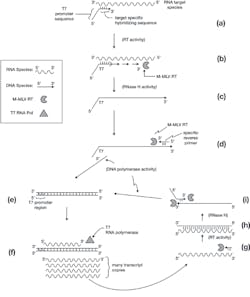
Figure 1. Transcription mediated amplification
a) A nucleic acids sample containing an RNA target of interest (with known sequence regions) is mixed with a synthetic DNA primer, much like a PCR primer. In this case, however, the primer has two sections. The 5′ section contains an RNA transcriptional promoter sequence (usually, and as labeled here, a promoter as recognized by the commercially available purified RNA polymerase from bacteriophage T7). The 3′ (extendable) end of this primer is a sequence designed as complimentary to a region of the target RNA species. Upon the mixing of the primer and RNA in a suitable buffer at some moderate temperature (say 56°C), target specific annealing of the T7 tagged primer to its RNA partner occurs. Note that, unlike with PCR, since the target species was already single-stranded (the natural form of the RNA target), no high temperature denaturation step was required to allow primer access to its homologous pairing region. Like PCR, however, this pairing is driven by sequence homology, and if the correct matching target sequence is not present, the T7-tagged primer will not successfully anneal.
b) The buffer contains a reverse transcriptase (RT) enzyme—that is, an enzyme which uses an RNA template molecule to direct the polymerization of a DNA complementary strand. Like a standard DNA polymerase, it creates a nascent growing strand in the 5′ to 3′ direction, and it must start from a pre-existing, annealed, 3′ -OH of a primer sequence. Most commonly, the RT used here is the Moloney murine leukemia virus RT (M-MLV RT); in addition to its pure RT activities, it has other enzymatic functions which the technique will make use of. At this step, the M-MLV RT extends the annealed T7-tagged primer and creates a DNA complement of the RNA target species.
c) One of the M-MLV RT’s other activities is an “RNase H” function. The H stands for hybrid, and this enzymatic activity degrades away the RNA pair of annealed DNA-RNA hybrids. The product of this step is therefore a single-stranded DNA complement copy of the original RNA target.
d) As with PCR, the TMA method requires a second, sequence specific “reverse primer,” opposite in polarity and downstream of the T7-tagged primer. This anneals to the single-stranded DNA product of the previous step, providing a starting point for another of M-MLV RT’s activities: a classical DNA polymerase function.
e) Extension of this by M-MLV RT in its DNA polymerase mode completes creation of a double-stranded DNA copy of the original RNA target region. Note this incorporates the T7 promoter region at one end of the molecule.
f) The reaction mixture also contains a second enzyme—an RNA polymerase (in this case, T7 RNA polymerase). This enzyme recognizes the double-stranded T7 promoter region of the DNA molecule in (e), and drives the production of large numbers of RNA transcript copies of the region. Note that these are opposite in polarity to the original RNA target molecule, and that they include the entire region between the target-specific portions of the two primers, but not the T7 promoter tag.
g) Each of the transcript molecules can now be annealed to by the reverse primer, providing a starting point for M-MLV RT to reverse transcribe and make a DNA copy of the transcript.
h) M-MLV RT’s RNase H activity now degrades the RNA component of this RNA/DNA hybrid double-stranded molecule.
i) This leaves a single-stranded DNA target copy, which can be annealed to by the T7-tagged primer. This provides two places for M-MLV RT to engage its DNA polymerase function: one at the 3′ end of the T7-tagged primer (generating the second “upper” strand of the figure), and one at the 3′ end of the DNA strand from (h), using the T7 promoter tag region of the primer as template. Together, these two strand extensions completely regenerate the dsDNA molecule of (e) and create an amplification cycle.
Since each dsDNA molecule (e) can drive the transcription of large numbers of RNA transcripts (f) and each of these can in turn create a new dsDNA (e), the system effectively amplifies the target region. In a 30-minute TMA reaction, a single target RNA template can be amplified to as many as 10e9 dsDNA copies. This amplification efficiency approaches that of PCR, and with similar specificity as defined by the need for two primers to anneal to matching target sequences. Unlike PCR however, TMA occurs with all steps at a single temperature (an “isothermal” process), removing the need for programmable thermal cycling instrumentation.
Downsides of TMA include that it is challenging to amplify and detect double-stranded DNA targets, as their double-stranded nature precludes the initial primer annealing step needed to start the process. This can be handled by an initial thermal denaturing and annealing step between the dsDNA target and the T7-tagged primer, followed by addition of the thermally labile M-MLV RT and T7 polymerase; however, this is less convenient than the direct method applicable to RNA target molecules. The need for two enzymes with very particular functions also reduces the flexibility of the method as compared to classical PCR, where a wide spectrum of modified DNA polymerases (with varying attributes such as resistance to particular reaction inhibitors) is available to “fine tune” an assay to particular sample types.
Detection of a positive signal in a TMA reaction is based on the detection of the dsDNA product molecule after completion of the amplification stage. While a number of approaches to this could be possible, similar to those employed in endpoint PCR product detection, most commercial applications of the method employ a probe-based chemiluminescent approach. In these, a chemically labeled synthetic DNA reporter probe is present; this probe is complementary to the possible amplified sequence. Where no product is formed, the probe cannot anneal and is chemically degraded by the detection procedure without emitting a signal. If a product it can anneal to is formed—that is, a successful TMA amplification has occurred—this annealing protects the reporter probe from degradation, and the probe emits a light signal detected by a luminometer. As with classical PCR, this use of a probe based detection method adds a further level of specificity to the detection process, as both primers and reporter probe must successfully hybridize to get end signal. Two forms of probe label chemistry may be used, one which emits a short burst of light immediately on substrate contact (“flashers”) and one which emits a slow steady glow over a longer period after substrate exposure (“glowers”). By incorporating both of these in a reaction and having the luminometer monitor both a short time period and a long time period, it is possible to multiplex TMA reactions to two targets and allow for use of internal controls.
Overall, TMA is a simple, rapid, and effective method and can be the method of choice for qualitative detection of some RNA targets such as viruses. Today’s molecular laboratorian can expect to encounter NASBA and specifically TMA-based methodologies in a number of commercial assays in common use.
About the Author

John Brunstein, PhD
is a member of the MLO Editorial Advisory Board. He serves as President and Chief Science Officer for British Columbia-based PathoID, Inc., which provides consulting for development and validation of molecular assays.