New applications of flow cytometry in the clinical lab
Beginning inside
The earliest clinically related application of cytometry focused on measuring the quantity of DNA in individual cells. Single cell measurements of DNA content in human tumors, initially performed using imaging techniques, were heavily influenced by early work at Los Alamos National Laboratory that focused on the development of flow cytometry technologies for cell cycle analysis,1 studying the effects of radiation on DNA, and, as a natural extension, the impact of then emerging cancer chemotherapies on the cell cycle.2 In the 1970s, this led to the widespread use of flow (and image) cytometry for measuring the DNA content (DNA ploidy) of human tumors. After Hedley, et al3 developed a technique to measure DNA content from nuclei isolated from paraffin-embedded tissues, DNA ploidy came to be widely utilized in clinical medicine. In a number of human tumors, an abnormal amount of DNA (DNA aneuploidy) in tumor cells (or nuclei) was believed to provide a marker for poorer prognosis.
In the early 1990s, as clinical DNA content measurements were increasingly being used, some members of the clinical flow community became concerned by the lack of consensus on techniques, standards and rules for the interpretation of DNA content data.4 The lack of standardization was reflected in an article in the Journal of Clinical Oncology reviewing the status of biomarkers for breast and colorectal cancers5 which stated that “present data are insufficient to recommend DNA Ploidy or S-Phase for management (breast or colorectal) or for prognosis (breast Ca).” Since this journal had a much higher impact on clinical practice than articles in cytometry-focused journals, oncologists ceased using DNA content measurements.
Going outside
In the early 1980s, emerging research into AIDS demonstrated the significant loss in CD4+ lymphocytes in peripheral circulation in individuals having other clinical indicators then used to diagnose AIDS. This observation rapidly evolved into an application for CD4 enumeration, which still represents a major clinical use of cytometry.
Within the same time period, technical advances in reagents (the introduction of two-color flow, which first used FITC and Texas Red6 and then exploded with the use of phycobiliproteins7) and in instrumentation (including the introduction of two-laser cytometers capable of simultaneous measurement of three, and then four, independent fluorochromes) were paralleled by clinical research that began to unravel the complexity of normal white blood cell differentiation.8
At the same time, translational clinical research was identifying cell surface markers (and later cytoplasmic markers such as TdT) that are useful in identifying hematoplastic neoplasias. With the increasing understanding of the complexity of normal hematopoiesis and of the characteristic differences in some hematopoietic malignancies achieved using phenotypic analyses, applications for clinical flow cytometry, such as the characterization of leukemias and lymphomas and the detection of minimal residual disease, began to evolve.
Back inside?
Although clinical immunophenotyping uses cytoplasmic markers (TdT, light chains, CD3, etc.), at present the majority of markers used are found on the cell surface. The current trend is to simultaneously measure six or more markers in one sample, providing more correlations for the expression of multiple markers on individual neoplastic or reactive cells.
In the late 1990s, research studies indicated that individuals with chronic lymphocytic leukemia (CLL) who progress to aggressive disease are more likely to have immunoglobulin heavy chain V-region genes (IgVH) in the germline configuration (as opposed to mature B-cells with rearranged heavy chain V-region genes).9 Subsequently, several groups demonstrated a relationship for intracellular ZAP-70 protein expression with either germline IgVH or with disease progression. Within the next three years, more than 50 papers were published on ZAP-70 expression and CLL.
Despite early reports, there is now little enthusiasm in the oncology community for the use of a flow-based ZAP-70 assay for CLL patients for two important reasons: first, there are no therapeutic regimens useful for specifically treating aggressive disease in CLL; and second, the clinical flow community was not successful in developing and implementing techniques, standards, and rules for the interpretation of ZAP-70 analysis by flow. Which negative and/or positive control to use for flow cytometry-based ZAP-70 expression was an issue.
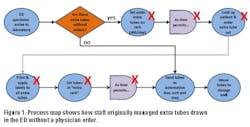
A single edition of the clinical cytometry journal Cytometry B,10 which reviewed most of the different flow-based assays and ZAP-70 controls, contained eight different approaches, including the use of internal T-cell and/or B-cell controls, the use of external B-cell (negative) controls, fluorescent bead standards (MESF), and isotype and isoclonic controls. As shown in Figure 1, whole blood samples provide both negative (normal B-) and positive (T-) cells that provide internal controls (identified using cell surface markers) for ZAP-70 measurement in individual samples. With this rich choice of potentially useful controls, the clinical flow community failed to develop a useful or timely consensus, a result similar to what happened 20 years earlier with clinical DNA content analysis.
ZAP-70 may represent a harbinger of things to come for clinical flow, in that it uses both cell surface and intracellular markers. ZAP-70 is normally part of a signal transduction network that links T-cell receptor MHC-antigen complex binding to specific downstream signaling pathways. Since the initial publication showing that specific signaling pathways can be monitored in whole blood samples using flow cytometry,11 numerous research publications have extended this observation. For example, bone marrow samples from acute myeloid leukemia (AML) patients show different signaling patterns than those seen in normal marrow.12-14 These studies used a combination of cell surface markers to identify specific cell populations (e.g., CD34 plus CD117 to identify myeloid progenitor cell populations) in conjunction with antibodies to identify specific intracellular signaling molecule activation.
In the majority of bone marrows studied, AML patients demonstrated a distinctly different signaling pattern from that seen in normal marrow. These types of data suggest that “Signaling Cytometry[I1]” may be useful in identifying leukemic cell populations, even when they are present in low numbers (potentially useful in minimal residual disease or MRD). What is perhaps more exciting is the ability to use this technology to identify the impact of drugs targeting specific signaling pathways. The publication by Chow et al11 showed that ex vivo treatment of normal whole blood with different MAP kinase inhibitors could be monitored using flow cytometry, and that the inhibition of a specific signaling protein (kinase) could be identified. “Signaling Cytometry” has also been used to monitor the impact of therapies on specific signaling pathways and has been shown to predict responses in individual patients.15 Thus, it may be possible to test a sample for responses to particular drugs ex vivo before treatment, and use similar flow cytometry analyses to monitor patients’ responses during therapy.
Back to the future?
In the 1990s, there was the perhaps naive concept that sequencing the human genome would provide a roadmap to normal cell function, and identify genetic changes that are related to disease (“one gene, one protein”). We now understand that regulation of cell functions is significantly more complex, and includes post-transcriptional protein modifications. Signal transduction pathways (which are the result of phosphorylation of specific amino acids on specific target proteins) represent only one important type of protein modification.
While the potential to monitor signaling pathways in real time16 offers a new and unique application for clinical flow cytometry, translational research is identifying other areas in which measurement of post-transcriptional protein modifications may prove useful in patient management. One rapidly growing research area is epigenetic modifications of DNA and proteins. Histone protein modifications (e.g., methylation, ubiquination, or acetylation) are involved in the regulation of cellular differentiation and proliferation, and aberrant expression has been associated with a number of disease states, including cancer, neuropsychiatric disorders, inflammation, and metabolic disorders.17
Some of these modifications have been measured using flow cytometry,18 and it is possible that as greater understanding is gained of epigenetic mechanisms and their role in human diseases, this area will provide a rich source of new, clinically useful flow cytometry-based assays. That said, the usefulness of these new “proteomic” focused flow assays will again depend on the adoption of standardized methodologies, standards, controls, and common methods of data analysis and interpretation. Internal cell populations (as with the ZAP-70 assay in CLL) and stabilized positive and negative reference cells could provide important standards and controls for these types of clinical assays. Without these, we face the prospect of expending much energy and money on developing assays that clinicians find uninformative or clinically irrelevant with DNA content and ZAP-70 flow assays.
T. Vincent Shankey, PhD, completed his doctoral work at the University of Florida School of Medicine in Immuonology and Medical Microbiology, and was a post-doctoral research fellow in the Department of Pathology at the University of Pennsylvania School of Medicine. He has previously worked in academia (Penn State University School of Medicine and Loyola University of Chicago Medical School) and has worked in a research group at Beckman Coulter, Inc., for the past 11 years.
References
- Tobey RA, Crissman HA, Kraemer PM. A method for comparing effects of different synchronizing protocols on mammalian cell cycle traverse. The traverse perturbation index. J Cell Biol. 1972;54:638-645.
- Tobey RA, Oka MS, Crissman HA. Differential effects of two chemotherapeutic agents, streptozotocin and chlorozotocin, on the mammalian cell cycle. Eur J Cancer. 1975;11:433-441.
- Hedley DW, Friedlander M, Taylor IW, et al. Method for analysis of cellular DNA content of paraffin-embedded pathological material using flow cytometry. J Histochem Cytochem. 1983;11:1333-1335.
- Shankey TV, Rabinovitch PS, Bagwell B, et al. Guidelines for implementation of clinical DNA cytometry. Cytometry. 1993;14:472-477.
- Clinical practice guidelines for the use of tumor markers in breast and colorectal cancer. J Clin Oncol. 1996;14:2843-2877.
- Herzenberg LA, Hayakawa K, Hardy RR, et al. Molecular, cellular, and systemic mechanisms for regulating IgCH expression. Immu. Rev.1982;67:5-31.
- Oi VT, Glazer AN, Stryer L. Fluorescent phycobiliprotein conjugates for analyses of cells and molecules. J Cell Biol. 1982;93:981-986.
- Civin CI, Loken MR. Cell surface antigen expression on human marrow: dissection of hematopoietic development using monoclonal antibodies and multi-parameter flow cytometry. Int J Cell Cloning. 1987;5:267-288.
- Hamblin TJ, Davis Z, Gardner A, et al. Unmutated IgV(H) genes are associated with a more aggressive form of chronic lymphocytic leukemia. Blood. 1999;94:1848-1854.
- Marti G, Orfao A. ZAP-70 in CLL: towards standardization of a biomarker for patient management. Cytometry B Clinical Cytometry. 2006;70;197-314.
- Chow S, Patel H, Hedley DW. Measurement of MAP kinase by flow cytometry using phosphor-specific antibodies to MEK and ERK: potential for pharmacodynamics monitoring of signal transduction inhibitors. Cytometry. 2001;46:72-78.
- Woost PG, Solchaga LA, Myerson HJ, et al. High-resolution kinetics of cytokine signaling in human CD34/CD117-positive cells in unfractionated bone marrow. Blood. 2011;117:e131-e141.
- Marvin J, Swaminathan S, Kraker G, et al. Normal bone marrow signal-transduction profiles: a requisite for enhanced detection of signaling dysregulation in AML. Blood. 2011; 117:e120-e130.
- Gibbs KD, Gilbert PM, Sachs K, et al. Single-cell phosphor-specific flow cytometric analysis demonstrated biochemical and functional heterogeneity in human hematopoietic stem and progenitor compartments. Blood. 2011;117:4226-4233.
- Brandwein JM, Hedley DW, Chow S, et al. A phase I/II study of imatinib plus reinduction therapy for c-kit-positive relapsed/refractory acute myeloid leukemia: inhibition of Akt activation correlates with complete response. Leukemia. 2011;25:945-952.
- Hedley DW, Chow S, Goolsby C, et al. Pharmacodynamic monitoring of molecular-targeted agents in the peripheral blood of leukemia patients using flow cytometry. Toxicol Pathol. 2008;36:133-139.
- Arrowsmith CH, Bountra C, Fish PV, et al. Epigenetic protein families: a new frontier for drug discovery. Nature Rev Drug Discovery. 2012;11:384-400.
- Chung EJ, Lee S, Sausville EA, et al. Histone deacetylase inhibitor pharmacodynamics analysis by multiparameter flow cytometry. An Clin Lab Sci. 2005;35:397-406.
- Shankey TV, Forman M, Scibelli P, et al. An optimized whole blood method for flow cytometric measurement of ZAP-70 protein expression in chronic lymphocytic leukemia. Cytometry B Clinical Cytometry. 2006;70:259-269.