Revealing the therapeutic potential of novel drugs for HPV-positive head and neck cancer
Human papillomavirus (HPV) is a viral infection that has well-known oncogenic effects. The virus is predominantly associated with the onset of cervical cancer, but its role in other malignancies has more recently been established. Effective antiviral therapies for HPV infection are lacking, and exciting new treatments may lie in the use of novel epigenetic drugs — bromodomain and extra-terminal domain inhibitors (BETis) — in combination with radiotherapy. This article describes recent research into the relevance and specificity of bromodomain-driven transcription in HPV-induced head and neck cancers and outlines the intriguing findings that have been revealed through next generation sequencing (NGS).
The HPV family includes more than 170 different viral strains that infect stratified epithelium, the protective layer of tissue covering all internal and external surfaces of the body.1 Approximately 18 percent of all human cancers have a viral etiology,2 and HPV has been identified as a key player in malignancies such as cervical cancer, anogenital tumors, and a subset of head and neck carcinomas. In particular, head and neck squamous cell carcinomas (HNSCCs) — HPV16-derived malignancies that develop from the mucosal epithelium in the oral cavity, pharynx and larynx — are responsible for more than 650,000 new cases of cancer annually, in addition to over 350,000 deaths.1 The frequency and severity of these carcinomas have led to a research surge in this field in recent years, driven by advancing technologies such as automated NGS library preparation. This has improved access to sequencing, leading to the identification of genomic and epigenetic factors involved in cancer progression, as well as the characterization of viral integration sites into the human genome.2
The biology of HPV infection in head and neck cancer
All papillomaviruses have a circular, double-stranded DNA genome of approximately eight kilobases that encodes eight or nine open reading frames (ORFs) — genes enabling viral genome replication and particle assembly during infection.3 These ORFs are organized and named based on the timing of expression; genes are either expressed early or late in the viral life cycle. Of the proteins encoded by the early expression genes, E1 and E2 are responsible for replication of the viral genome, while E4 to E7 are accessory proteins that coordinate viral genome amplification and virulence. The late expression genes — known as L1 and L2 — encode two coat proteins that make up the viral shell and enable its entry into host cells.3,4
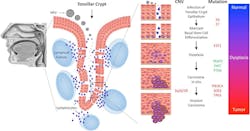
To establish an infection, papillomavirus virion must gain access to the epithelial basal layer of cells in the tonsillar crypt, entering the dividing basal keratinocytes and delivering their DNA into the nucleus of the cells (Figure 1).5,6 Viral DNA normally exists separately to the host genome in the nucleus as an episome, employing enzymes in the host cell to replicate its genome, along with host chromosomes, in a carefully regulated manner.3,4 Tight regulation of the replication process keeps copy numbers of the viral DNA low, helping the virus to evade the host immune system. However, in oncogenic HPV genotypes, E6 and E7 oncoproteins enable the integration of HPV into the host genome, dysregulating the normal cell cycle of keratinocytes in favor of viral replication. These oncoproteins control expression of p53 and Rb tumor suppressor genes, boosting cellular proliferation while preventing normal repair of chance mutations in the cellular genome, and driving HPV-related carcinogenesis.3
An epigenetic target
The initiation and progression of HNSCC and other cancers occur due to both genetic changes, such as genome instability and mutation, and dysregulation of the epigenome.7,8 Reversible epigenetic modifications —including DNA methylation, histone modification, chromatin remodeling and noncoding RNAs — regulate various cellular responses, and disruptions to these processes can lead to heightened expression of oncogenes, playing an important role in tumorigenesis. A classic example of this is the acetylation of lysine residues, a process mediated by histone acetyltransferases (HATs), at enhancer regions of oncogenes, such as c-MYC, BCL2 and CDK6. Histone acetylation can be recognized by proteins carrying bromodomains (BRDs) and, in turn, these epigenetic readers can recruit chromatin complexes to modulate transcription and activate the expression of oncogenes – such as E6 and E7 – that play an important role in cancer initiation and progression (Figure 2).9,10
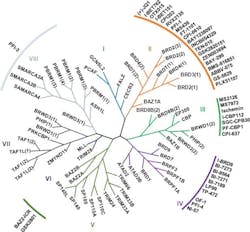
The human genome encodes 46 diverse proteins containing BRDs, which are structurally clustered into eight distinct subfamilies.10 The second subfamily — consisting of BRD2, BRD3, BRD4 and BRDT — is known as bromodomain and extra-terminal domain (BET) proteins. This group of epigenetic readers has recently emerged as a promising target for cancer therapeutics, with the development of small molecule inhibitors to prevent the transcription of key genes involved in cancer pathogenesis.
NGS developments
Understanding the potential of BET inhibitors to mediate cancer requires in-depth knowledge of the role of BET oncoproteins, and this has been the focus of multiple genomic and epigenetic studies, as well as more than 39 clinical trials around the world. For example, researchers in the Department of Human Oncology at the University of Wisconsin-Madison have conducted extensive investigations into the specific functions of these oncoproteins. The team characterized a panel of HPV-infected cell lines for their full integration status and examined the distinct roles of BRD2 and BRD4 in regulating viral-host transcription machinery. Determining the downstream effects of these proteins on DNA repair, cell cycle control, and oncogenic signaling pathways required a combination of NGS and biochemistry techniques.
DNA sequencing technologies have evolved rapidly over the last few decades, and NGS enables the analysis of millions to billions of DNA or RNA sequences in parallel.11 This offers many advantages over traditional sequencing methods — including higher throughput, reduced costs, and easier data interpretation — but also requires advanced equipment and information technology to handle upstream and downstream processes, such as library preparation and analysis of the large amounts of data that are generated. Researchers in the Department of Human Oncology have therefore introduced a number of new technologies into their lab, including a robust and reliable benchtop library preparation solution that streamlines all the steps required to prepare nucleic acid libraries. The introduction of this equipment reduces the time and effort required for sample preparation, allowing researchers to focus on understanding the biological underpinnings of HPV-induced HNSCC. Automated library preparation platforms, such as Tecan’s MagicPrep™ NGS system, has improved the consistency of data generated by the laboratory, and enhanced the researchers’ confidence in the accuracy of the transcriptional data that they produce.
Improving treatment for HPV-related cancers
The University of Wisconsin-Madison team has focused its efforts on answering several questions related to HPV HNSCC, such as why oncogenic HPV genotypes integrate into the genome, whether BRD4 coregulates viral and host genes in the presence of a BET inhibitor, and whether the antitumorigenic effects of BTE inhibitors occur independently or in the presence of radiation. The results are yet to be published, but they could potentially lead to improvements to conventional HPV treatments, which combine cisplatin administration and radiation therapy. Although cisplatin is an effective chemotherapeutical agent, its administration often leads to multiple side effects — including nephrotoxicity12 — and the onset of additional comorbidities. These adverse effects occur because cisplatin is a non-specific DNA cross-linking agent; it binds randomly to genetic material and activates multiple signaling pathways in host cells, resulting in excessive inflammation, oxidative stress, and damage in healthy cells.
Conclusion
Researchers at the University of Wisconsin-Madison plan to continue their research into BET inhibitors, using the new automated library preparation system to accelerate NGS workflows and enhance the reproducibility of their results. They are currently exploring new bromodomain-specific inhibitors targeted at BRD1 or BRD2 and optimizing the effectiveness of this new class of non-toxic anticancer drugs. In the long term, BET inhibition holds significant clinical potential as part of combination treatment regimens for HPV HNSCC, and for a variety of other HPV-related cancers.
REFERENCES
- Sabatini ME, Chiocca S. Human papillomavirus as a driver of head and neck cancers. Br J Cancer. 2020;122(3):306-314. doi:10.1038/s41416-019-0602-7.
- Tuna M, Amos CI. Next generation sequencing and its applications in HPV-associated cancers. Oncotarget. 2017;31;8(5):8877-8889. doi:10.18632/oncotarget.12830.
- Faraji F, Zaidi M, Fakhry C, Gaykalova DA. Molecular mechanisms of human papillomavirus-related carcinogenesis in head and neck cancer. Microbes Infect. 2017;19(9-10):464-475. doi:10.1016/j.micinf.2017.06.001.
- Doorbar J. Molecular biology of human papillomavirus infection and cervical cancer. Clin Sci (Lond). 2006;110(5):525-41. doi:10.1042/CS20050369.
- Aksoy P, Gottschalk EY, Meneses PI. HPV entry into cells. Mutat Res Rev Mutat Res. 2017;772:13-22. doi:10.1016/j.mrrev.2016.09.004.
- Faraji F, Zaidi M, Fakhry C, Gaykalova DA. Molecular mechanisms of human papillomavirus-related carcinogenesis in head and neck cancer. Microbes Infect. 2017;19(9-10):464-475. doi:10.1016/j.micinf.2017.06.001.
- Kanwal R, Gupta K, Gupta S. Cancer epigenetics: an introduction. Methods Mol Biol. 2015;1238:3-25. doi:10.1007/978-1-4939-1804-1_1.
- Cheng Y, He C, Wang M, et al. Targeting epigenetic regulators for cancer therapy: Mechanisms and advances in clinical trials. Signal Transduct Target Ther. 2019;17;4:62. doi:10.1038/s41392-019-0095-0.
- Suarez-Alvarez B, Rodriguez RM, Ruiz-Ortega M, Lopez-Larrea C. BET Proteins: An Approach to Future Therapies in Transplantation. Am J Transplant. 2017;17(9):2254-2262. doi:10.1111/ajt.14221.
- Pérez-Salvia M, Esteller M. Bromodomain inhibitors and cancer therapy: From structures to applications. Epigenetics. 2017;4;12(5):323-339. doi:10.1080/15592294.2016.1265710.
- Hess JF, Kohl TA, Kotrová M, et al. Library preparation for Next generation sequencing: A review of automation strategies. Biotechnol Adv. 2020;41:107537. doi:10.1016/j.biotechadv.2020.107537.
- Sun L, Liu J, Yuan Y, Zhang X, Dong Z. Protective effect of the BET protein inhibitor JQ1 in cisplatin-induced nephrotoxicity. Am J Physiol Renal Physiol. 2018;1;315(3):F469-F478. doi:10.1152/ajprenal.00527.2017.
About the Author

Ashesh Saraiya, PhD
is the Associate Director of Product Management at Tecan Genomics. He completed his PhD in molecular biology at Wayne State University, specializing in ribosomal RNA genetics, and followed this up with postdoctoral work examining miRNA and gene translation in protozoan parasites at University of California, San Francisco. Ashesh began his Tecan career as a Field Application Scientist, before moving to product management, where he continues to leverage his experience to help develop innovative solutions to address the challenges associated with NGS sample preparation.