Copy number variation detection with next generation sequencing data: the impact on pharmacogenetics
Copy number variations, changes in the frequency of particular genetic sequences, are a critical element for pharmacogenomics, as they can have a significant impact on human phenotypes, including links to a variety of diseases such as cancer,1 schizophrenia,2 and autism.3,4 Variations in copy number can range from relatively small regions, such as a single gene or just part of a gene, to larger regions, such as an entire chromosome.
A variety of technologies have been applied to the detection of copy number variations. These include microarray methods, including SNP arrays and array comparative genomic hybridization (aCGH), quantitative PCR (qPCR), and quantitative fluorescent PCR (qfPCR). The aCGH method is based on measuring the co-hybridization of fluorescently labeled sample and control DNA. The fluorescent intensity relative to the control is measured to determine the copy number. Fluorescent tags are also used with the qPCR and qfPCR methods. With the qPCR method, the DNA is quantified real-time during the PCR process. The PCR products are labeled with fluorescent tags and the increased fluorescence during the exponential phase of PCR is measured. With qfPCR, different colored fluorescent tags can be used for different primers and the amplified product can be evaluated, with two peaks expected for the normal heterozygous case. For example, an additional peak or increased height of one of the peaks is observed in the case of trisomy, where three copies are present. An example of trisomy detected by qfPCR is shown in Figure 1.
Next generation sequencing (NGS) technologies, also known as high-throughput sequencing, have had a great impact on biological research5 and clinical diagnostics.6 These sequencing systems can be useful for detecting copy number variants. They are capable of producing a large amount of sequence data at a reduced cost compared to previous sequencing methods, and the same sequence data can be applied to both CNV analysis and other applications such as SNP and indel detection. NGS techniques can be utilized to evaluate copy number variations based on targeted gene panels, whole exome sequencing, and whole genome sequencing.
To perform copy number variation analysis, the coverage depth (quantity of sequence reads) for targeted regions between a sample and control can be compared. For a particular region, the ratio of the coverage of the sample to the total coverage (sample + control) can indicate the CNV status for that region. For a normal copy number compared to the control the ratio would be expected to be 0.5. This is based on two copies for both the sample and the control, which gives a ratio of 2/4 (sample/total), or 0.5. For a heterozygous insertion, where the sample contains three copies and the control two, the ratio is expected to be 0.6 (3/5).For a heterozygous deletion, where the sample contains one copy and the control two, the ratio is expected to be 0.33 (1/3).
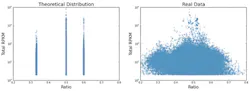
However, when working with NGS data, unfortunately the copy number is not the only factor that determines the coverage. Noise in the data means that there will generally be a more broad dispersion of ratios, which makes effectively determining copy number challenging. Figure 2 shows a comparison of the theoretical and actual distribution of coverage ratios. For this reason, additional analysis is needed to evaluate the coverage ratios for determination of copy number. A beta-binomial model can be fit to the coverage ratios to model the noise. Generally the noise decreases with increased coverage depth. This enables the use of generating a statistical model for coverage and copy number. Probabilities can be calculated based on the coverage ratios and the dispersion model. A Hidden Markov Model (HMM), a statistical modeling method, can be used to determine the copy number.7,8 The Hidden Markov Model allows noise (small regions with extreme coverage ratios) to be ignored, while longer regions with less extreme ratios can be correctly determined as CNVs.
Figure 3 shows the CNV results with this method for a whole exome sequencing dataset from a tumor sample compared with a matched control sample. Data in gray (generally near the ratio of 0.5) represents normal regions, while data in red (near the ratio of 0.33) represents deletions and data in green (near the ratio of 0.6) represents insertions.
A further application of copy number variation that is in the early stages of application for next generation sequencing is noninvasive prenatal testing (NIPT), which is used to test for chromosomal abnormalities. Noninvasive methods pose less risk to mother and fetus than common invasive methods such as amniocentesis and chorionic villus sampling (CVS). NIPT is based on using maternal blood plasma, which contains fetal DNA in the form of cell-free DNA (cfDNA).9 Novel approaches for CNV analysis are required to work with this type of data, as the sequencing results represent both maternal and fetal DNA.
References
- Shlien A, Malkin D. Copy number variations and cancer susceptibility. Curr Opin Oncol. 2010;22:55–63
- Mulle JG, Dodd AF, McGrath JA, et al. Microdeletions of 3q29 confer high risk for schizophrenia. Am J Hum Genet. 2010;87(2):229-236.
- Sebat J, Lakshmi B, Malhotra D, et al. Strong association of de novo copy number mutations with autism. Science. 2007;316(5823):445–449
- Kusenda M, Sebat J. The role of rare structural variants in the genetics of autism spectrum disorders. Cytogenet Genome Res. 2008;123(1–4):36–43.
- Schuster SC. Next-generation sequencing transforms today’s biology. Nat Methods. 2008;5:16-18.
- Sikkema-Raddatz B, Johansson LF, de Boer EN, et al. Targeted next-generation sequencing can replace Sanger sequencing in clinical diagnostics. Hum Mutat. 2013;34(7):1035-1042
- Fromer M, Moran JL, Chambert K, et al. Discovery and statistical genotyping of copy-number variation from whole-exome sequencing depth. Am J Hum Genet. 2012; 91(4):597–607.
- Simpson JT, McIntyre RE, Adams DJ, Durbin R. Copy number variant detection in inbred strains from short read sequence data. Bioinformatics. (2010);26(4): 565-567.
- Swanson A, Sehnert AJ, Bhatt S. Non-invasive prenatal testing: technologies, clinical assays and implementation strategies for women’s healthcare practitioners. Curr Genet Med Rep. 2013;1(2):113–121.