Improving the molecular diagnosis and treatment of epilepsy with complex genetic testing
Epilepsy is a highly complex disease and is one of the most common neurological conditions worldwide. Some 2,500 years after Hippocrates observed a hereditary tendency for epilepsy, researchers discovered the first causative gene, neuronal nicotinic acetylcholine receptor alpha 4 subunit (CHRNA4), in 1995.1,2 With the development and implementation of new genomic profiling technologies, the discovery of epilepsy-associated genes has increased rapidly in the past five years. It is currently estimated that 70 percent to 80 percent of epilepsy cases have a genetic component.3 Incorporating comprehensive genetic testing for epilepsy into clinical practice has enabled physicians to provide a proper diagnosis for many patients, resulting in a better understanding of prognosis, family counseling, and targeted treatment options.
Due to both the diverse spectrum of causative mutations, and the large number of genes and genomic regions associated with epilepsy, several molecular technologies are required to perform the most comprehensive genetic testing and produce the highest diagnostic yield. Genome-wide chromosomal microarray analysis (CMA) to detect copy number variants (CNVs) is generally the first line of testing for patients with unexplained epilepsy with co-morbid neurodevelopmental features.4 Current high-density oligonucleotide microarrays enable the detection of both large scale CNVs and small exon-level deletions and duplications in disease-associated genes. The diagnostic yield with CMA for individuals with epilepsy in association with intellectual disability or an autism spectrum disorder is estimated to be ~15 percent to 20 percent.5 CMA is also an appropriate first line test for individuals presenting with infantile spasms, with a diagnostic yield estimated at ~11 percent.6 Target enrichment and next-generation sequencing (NGS) technologies, such as pre-specified candidate gene panels and whole exome sequencing, have revolutionized the field of epilepsy genetics during the last five years. These tests are used routinely in the clinic today and offer clinicians a variety of options depending on the patient’s phenotype.
The cost effectiveness and availability of target enrichment and NGS technologies have resulted in a multitude of commercial laboratories offering a wide range of epilepsy testing options. It is important for clinicians to realize that all tests are not created equal and that the detection rate will vary depending on not only the gene content of the panels ordered, but also the sensitivity and specificity of the technology and bioinformatics used for testing. These factors all impact the quality of the test.
For example, labs using just NGS for testing will miss large repeat expansions, like the dodecamer repeat expansion within the 5’-untranslated region of the cystatin B (CSTB) gene, which is responsible for the vast majority of the Unverricht-Lundborg type of progressive myoclonus epilepsy cases.7 In addition, labs detecting mutations and CNVs utilizing NGS should use a secondary technology such as Sanger sequencing and microarray analysis or qPCR, respectively, in order to eliminate the potential of reporting false positives. Tests that are marketed as having a zero false positive rate using NGS data alone generally do not utilize a very sensitive bioinformatics pipeline, resulting in a higher rate of false negatives for mosaic mutations, indels, and mutations in highly complex genomic regions.
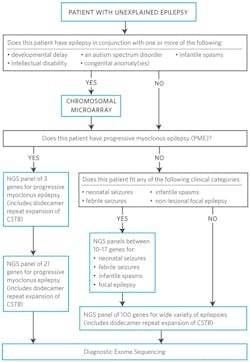
Although NGS provides a cost-effective method to sequence a multitude of genes concurrently, clinicians may want to consider ordering smaller phenotype-associated panels initially, with the option to reflex to a more comprehensive panel (Figure 1). This method provides clinicians the ability to target the genes most likely to be causing their patient’s epilepsy, minimize turnaround time (TAT), and limit reporting to only those genes that are highly characterized in the epilepsy phenotype of interest. With such a large number of genes being reported in association with epilepsy, most with very low rates of mutation detection, it is not surprising that commercial labs vary widely in the number of genes offered for nonspecific epilepsy presentations. For example, a recent publication reviewed epilepsy panels from seven different laboratories, and the number of genes analyzed varied from 70 to 465 genes.8 Some tests included genes with very little or no supporting evidence associating them with epilepsy.
Importantly, the more genes included on a panel, the more complex the test report and the longer the TAT, as the number of variants of uncertain significance increases. Clinicians should consider ordering exome sequencing once they have exhausted targeted panel options, or if they have a reason to be looking to interrogate more than ~200 genes at once.
Exome sequencing, analyzing the coding regions of ~20,000 genes, has significantly contributed to the rapid discovery of epilepsy-associated genes and has shortened the “diagnostic odyssey” for a large number of previously undiagnosed patients. It is estimated that exome sequencing has a diagnostic yield of ~38 percent for epilepsy.9 Similar to gene panel testing, exome diagnostic yield is dependent on the test design and the quality of interpretation. For example, not all commercially available exome tests utilize the same gene content or quality coverage metrics such as the percent of targeted bases covered at 10X. Likewise, only one or two labs contain the experience and resources needed to report novel genetic etiologies, which can provide a diagnosis to an additional ~9 percent of epilepsy patients.9
The introduction of NGS and enhanced diagnostic testing options has enabled clinicians to better understand the complex genetic contribution to epilepsy and determine a causative diagnosis for patients, leading to an appropriate counseling and treatment plan. This was recently illustrated by research presented at the 2015 American Epilepsy Society Annual Meeting, which indicated that ~16 percent of epilepsy patients who underwent diagnostic sequencing harbored a mutation in a gene which had immediate treatment implications.10 As the number of clinicians who are comfortable utilizing complex genetic testing increases, the understanding of epilepsy genetics will increase rapidly, precipitating a paradigm shift in the diagnosis, management, and treatment of the disorder.
References
- Riggs AJ, Riggs JE. Epilepsy’s role in the historical differentiation of religion, magic, and science. Epilepsia. 2005;46:452-453.
- Steinlein OK, Mulley JC, Propping P, et al. A missense mutation in the neuronal nicotinic acetylcholine receptor alpha 4 subunit is associated with autosomal dominant nocturnal frontal lobe epilepsy. Nat Genet. 1995;11(2):201-203.
- Hildebrand MS, Dahl HH, Damiano JA, et al. Recent advances in the molecular genetics of epilepsy. J Med Genet. 2013;50:271-279.
- Miller DT, Adam MP, Aradhya S, et al. Consensus statement: chromosomal microarray is a first-tier clinical diagnostic test for individuals with developmental disabilities or congenital anomalies. Am J Hum Genet. 2010;86(5):749-764.
- Mefford HC. CNVs in Epilepsy. Curr Genet Med Rep. 2014;2(3):162-167.
- Wirrell EC, Shelhaas RA, Joshi C, et al. How should children with West syndrome be efficiently and accurately investigated? Results from the National Infantile Spasms Consortium. Epilepsia. 2015;56(4):617-625.
- Lafrenière RG, Rochefort DL, Chrétien N, et al. Unstable insertion in the 5’ flanking region of the cystatin B gene is the most common mutation in progressive myoclonus epilepsy type 1, EPM1. Nat Genet. 1997;15(3):298-302.
- Chambers C, Jansen LA, Dhamija R. Review of Commercially Available Epilepsy Genetic Panels. J Genet Couns. 2015. Epub ahead of print:
doi:10.1007/s10897-015-9906-9. - Helbig KL, Farwell KD, Powis Z, et al. Diagnostic exome sequencing provides a molecular diagnosis for a significant proportion of patients with epilepsy. Genet Med. 2015; In press.
- Demos M, Buerki SE, Guella I, et al. Targeted analysis of whole exome sequencing in early onset epilepsy. American Epilepsy Society. 2015; Abstract 1.311.