The technology and clinical applications of hybrid capture NGS
Clinicians are challenged with identifying variants of disease and determining appropriate treatment plans on a daily basis. The more relevant the information they have at their fingertips, the more likely a successful patient outcome. Nucleic acid testing (NAT) is a common method of detecting the presence of specific disease-associated bacteria and viruses, such as HIV, and is used in routine blood screening. Though it is inexpensive and reliable, NAT cannot always provide clinicians with all the information they need, however, which is where a more detailed analysis of samples becomes necessary.
Whole genome next generation sequencing (NGS) can provide a wealth of information to direct treatment. However, it has been a costly option for clinicians, particularly for large numbers of samples. A more cost-effective option is targeted NGS, in which genomic regions of interest are enriched prior to sequencing. Targeted NGS focuses resources on acquiring relevant data and enables multiplexing, so many patient samples can be analyzed simultaneously. As such, certain applications can benefit from targeted NGS, from analyzing mutations in cancer to viral integration sites and variants.
While targeted NGS can be performed cheaply via polymerase chain reaction (PCR), processing the numbers of samples common in many clinical research facilities at the required level of quality and consistency is more suited to the approach of hybrid capture. This technology uses labelled DNA probes complementary to the target region for enrichment from fragmented genomic samples. Obtaining high efficiency and accuracy, as well as minimizing cost, requires careful consideration of probe design.
Optimized oligo design
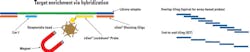
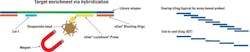
Oligonucleotides used for hybrid capture NGS can be designed based on a variety of technologies. One such technology, which is seeing increased use in clinical diagnostics, utilizes biotin-streptavidin interaction for enrichment. 5’ biotinylated probes are designed to hybridize to the target DNA, and isolated using streptavidin beads (Figure 1).
Instead of many shorter overlapping probes, this variation of the technology uses longer 120 bp probes designed for end-to-end tiling across the target region. This maintains a high capture efficiency while reducing the number of overall probes required, saving cost. In addition, this length of probe can tolerate several mismatches and still pull down the targets, affording the detection of novel variants.
The challenge with longer oligos, however, is that the coupling efficiency of each additional base affects the final yield of the full-length oligo. While obtaining a 99.6 percent efficiency results in 62 percent of products being full-length for 120-mers, losing just one percent efficiency reduces full-length yield to as little as 15 percent. Highly efficient synthesis in this variation of hybrid capture is coupled with biotinylation only of full-length product, thereby ensuring only the complete oligos are enriched, improving sequencing reliability.
This method of hybrid capture also employs blocking oligos for library adaptors to maximize targeting efficiency. These blocking oligos are especially important in reducing costs via multiplexing, as libraries carry unique barcodes to associate each read with the appropriate sample, which must be obscured during pull-down. Targeting may be improved by 60 percent through this method, resulting in lower cost and the generation of highly-focused clinically relevant data.
Quality control
High data quality, especially when searching for specific variants of a target, is essential in targeted sequencing panels.
Commercially synthesized hybrid capture oligos, such as the type described here, are produced individually, allowing more stringent and individualized QC. Specifically, electrospray ionization mass spectroscopy (ESI-MS) is used to confirm the absence of truncated species. Those samples found with truncated species can undergo re-synthesis and be QC’d again, saving waste.
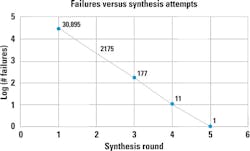
Interestingly, looking in more detail at quality-control data highlights the stochastic nature of synthesis failures (Figure 2). When analyzing an exome research panel of approximately 430,000 probes covering 39Mb, for every re-synthesis round, the number of failures decreased exponentially. This indicates that failure is random, rather than being sequence-specific, and as such, a re-synthesis approach could be applied to all oligos. This provides clinicians some assurance that their diagnostics results can be consistent regardless of target.
These data suggest that when oligos undergo only a single synthesis, as is the case with array-synthesis platforms, the probe population will have varying levels of truncation. This could lead to insufficient reproducibility of enrichment and therefore inconsistent NGS results.
By synthesizing, quality controlling, and normalizing every probe individually, the amount of each probe in the final pool can be determined. This benefits clinical applications through identical performance across batches. Moreover, commercial probe panels are available that conform to FDA 21 CFR Part 820 certification for medical devices (GMP), further ensuring quality for clinical applications.
For studies with large numbers of samples, this approach becomes highly cost-effective, especially as it allows a modular approach to probe panel design, with individual targets mixed to adapt the study for new regions of interest.
Targeted NGS in cancer
Acute myeloid leukemia (AML) is characterized by faulty differentiation of hematopoietic stem cells, deriving from mutations arising at cell division. In one case, on analysis of cells taken from bone marrow, a patient presented with four different subclones of AML cells with varying prevalence, as analyzed by targeted NGS. Following chemotherapy, none of the cancerous clones could be detected by standard blood analysis, although targeted NGS detected the clonal evolution of leukemic cells. The patient eventually relapsed as a result of the least prevalent subclone at diagnosis, which developed an additional mutation, proliferating in the absence of the others. NGS analysis allowed detection of this evolved clone before it was seen by standard blood characterization.
This example indicates how improved sequencing could increase the chances of detecting rare cancerous clones. In clinical diagnostics, increasing sequencing depth to 500x would bring the level of sensitivity required to detect clones down to five percent prevalence. Such a level of sequencing depth is only feasible through targeted NGS, and in these types of cases helps clinicians identify clues pointing to patient relapse.
Interestingly, it has also been indicated that diminishing the level of cancerous clones by chemotherapy to a level below five percent correlates with improved patient outcome,1 and this can be directly tracked with sensitive NGS.
Viral research
Targeted NGS also has utility in virology. For example, genomic integration of the human papillomavirus (HPV) can cause several cancers. Hybridization probes designed against the 8Kb HPV genome provide detection of the integration site. Sequencing the flanking regions can also provide insight into patterns in HPV integration and its role in cancer.
In addition to oncogenic viruses, infectious agents such as Ebola and Zika virus can be traced with targeted NGS. The ability of hybrid capture probes to tolerate several mismatches enables novel variants of rapidly mutating viruses to be detected. Different isolates can then be tracked to different geographic regions, potentially allowing the source of the outbreak to be identified.
Integrating NGS into the clinic
By bringing the cost of sequencing down, targeted NGS using hybrid capture probes can complement NAT strategies when more detailed information is needed.
Targeted NGS presents the latest innovation in sequencing technology, with cutting-edge probe and assay designs that reduce costs and improve reliability. Bringing the benefits of these new technologies into the clinic requires an interface between clinicians and genomics experts. As these technologies proliferate, in addition to growing in-house expertise, it is now becoming a popular option to look toward external partnerships for a complete solution, from bioinformatics to customized NGS panels.
REFERENCE
- Klco JM et al. Association between mutation clearance after induction therapy and outcomes in acute myeloid leukemia. JAMA. 2015;314(8): 811-822.
* Figure 1 – Oligo refers to xGen Blocking Oligo and Probe refers to xGen Lockdown Probe, both from IDT.
John Havens, PhD, serves as Vice President of Business Development at Coralville, Iowa-based Integrated DNA Technologies.