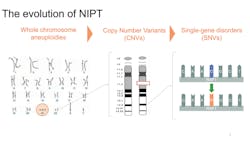
In the May 2015 issue of MLO [2015;47(5):14] we reviewed the evolution of prenatal genetic screening. The closing line of that article stated that “…rapid technological progress, particularly using SNPs, holds the promise of even greater improvements in test performance and safety.”1 Now, we revisit the status of prenatal screening for chromosome abnormalities and the continued advancements in this area.
Non-invasive prenatal testing or screening (NIPT/NIPS) using cell-free DNA (cfDNA) has been commercially available for over five years. A number of peer-reviewed journal articles have consistently demonstrated its superior positive predictive value for common trisomies (21, 13, and 18) compared to traditional maternal serum screening. Norton et al demonstrated a 79 percent sensitivity and 3.4 percent positive predictive value (PPV) for trisomy 21 using standard first-trimester screening in all patients. In comparison, cfDNA screening showed a 100 percent sensitivity and 81 percent PPV for trisomy 21 for all patients.2 Most important, a total of 11,994 women in that study were < 35 years of age, without additional risk factors for trisomy 21, and cfDNA showed 100 percent sensitivity and 76 percent PPV in that “low risk” group.
Both the American College of Obstetricians and Gynecologists (ACOG) and the American College of Medical Genetics (ACMG) have updated their screening statements to recommend that NIPT be made available to all pregnant woman, regardless of their prior risk for aneuploidy.3,4 However, approximately half of women with private insurance and essentially no women with Medicaid are financially covered to choose NIPT as a first-line screen.
The fact that NIPT maintains its high performance standards in average-risk women in turn prompts continued discussion to “rethink screening.” There are many conditions that are unrelated to maternal age, and usually without a family history. For what additional conditions beyond trisomies is it reasonable/responsible to offer screening to all pregnant women, if non-invasive screening is possible with acceptable sensitivity, specificity, and positive predictive values? How is SNP technology for NIPT uniquely positioned to provide sensitive and specific expansion of prenatal genetic screening?
A logical next step in prenatal genetic screening is to consider smaller genetic changes in a chromosome, called copy number variants (CNVs). CNVs are typically microdeletions or microduplications less than 10 Mb that are associated with clinically significant outcomes and are unrelated to maternal age. In 2012, Wapner et al reported that clinically relevant deletions and duplications were found in six percent of pregnancies with ultrasound anomalies and 1.7 percent of pregnancies without risk factors.5 However, unlike the common trisomies, deletions and duplications are not part of routine serum screening or NIPT. There are different approaches to CNV screening: targeted and whole genome.
Targeted screening
Targeted CNV screening entails choosing specific deletion or duplication sites with known clinical outcome, and providing risk assessment particular to that associated condition. For example, the most common microdeletion condition, 22q11.2 deletion syndrome, has a published incidence at birth of 1/2000, though this may be an underestimate. It is a well-described clinical entity with known chromosomal breakpoints and clear diagnostic testing parameters. Therefore, screening for 22q11.2 deletion syndrome is a natural expansion of NIPT.
ACOG and ACMG guidelines are clear with respect to the necessary laboratory conditions that need to be met in order to offer CNV screening. ACMG states that “Laboratory requisitions and pretest counseling information should specify the DR, SPEC, PPV, and NPV of each CNV screened.”4 This requires analytical and clinical validation for each deletion or duplication, as well as accurate estimates of population incidence of each condition. Currently, different NIPT laboratories offer a range of microdeletion syndromes, but their reports often do not follow these guidelines.
A recent publication on the clinical experience of SNP-based microdeletion testing addresses the issues of expanded screening and the need for transparent follow-up. This publication extends the initial reporting of SNP-based NIPT screening for 22q, and highlights outcome data for the remainder of the microdeletion panel currently offered (1p36 deletion, Angelman, Prader-Willi, and cri-du-chat). Performance improvements to this SNP-based testing resulted in a decrease in false positive test rate (0.07 percent for 22q) and an increase in PPV (44.2 percent for 22q; 31.7 percent combined for others). While some publications have questioned the expansion of NIPT into microdeletions due to concerns about positive screen rate and low detection rate, the targeted nature of SNP-based NIPT screening is shown to have a higher sensitivity than other NIPT methodologies.6
Whole genome
The concept of targeted CNV screening can be further expanded to other less common and less well-described microdeletions and microduplications, as well as rare autosomal trisomies (RATs), by looking at genomic information across all chromosomes. However, ACMG specifically recommends not screening for genome-wide CNVs, stating (among other reasons) that “If this level of information is desired, then diagnostic testing followed by CMA is recommended.”3 The clinical performance of such test options is not clinically well validated, and reporting does not follow test metric guidelines, as described earlier.
Single gene disorders
As the precision of non-invasive prenatal screening using SNP technology has narrowed in on CNVs, this screening can then focus even further to single gene disorders.
The first commercially available screening test for single gene disorders was launched in early 2017. This panel screens for single nucleotide variants (SNVs) in 30 genes responsible for a variety of genetic conditions. These genes can be generally categorized by clinical phenotype: Noonan spectrum, Craniosynostosis, Skeletal, and Syndromic disorders. Many of the conditions have no ultrasound findings early in pregnancy, are typically de novo, and may be associated with advanced paternal age. The combined incidence of these conditions in the general population is approximately one in 600.
Each of these genes are screened in the cfDNA of a pregnant woman using next-generation sequencing technology. Pathogenic or likely pathogenic variants are reported, with the recommendation for confirmation by diagnostic testing.
The view from here
Currently, prenatal screening options are typically limited to trisomies 13, 18, and 21, even though the general population incidence of other genetic conditions may be higher. Unfortunately, despite significant published data regarding the superior performance of NIPT over conventional screening, many women are denied access due to lack of insurance coverage.
Advances in SNP-based NIPT technology have allowed for the expansion of these prenatal genetic screening options for conditions unrelated to maternal age such as targeted microdeletions and single gene disorders. Indeed, the promise of greater improvements in non-invasive prenatal test performance has held true. It is exciting to consider the next advancements on the horizon. However, one critical fact remains unchanged: regardless of the kind of prenatal screening performed during pregnancy, the results are not diagnostic, and no irreversible decisions should be made on the basis of screening results. Confirmatory testing, either prenatally by chorionic villus sampling or amniocentesis or postnatally by peripheral blood draw, is required.
REFERENCES
- Gross S, McKanna T. The evolution of prenatal genetic screening. MLO. 2015;47(5):14. https://www.mlo-online.com/prenatal-screening-with-microarray-technology.php.
- Norton ME, Jacobsson B, Swamy GK, et al. NEJM. Cell-free DNA analysis for noninvasive examination of trisomy. N Engl J Med. 2015;372(17):1589-1597.
- American College of Obstetricians and Gynecologists. Practice Bulletin No. 163: Screening for fetal aneuploidy. Obstet Gynecol. 2016;127(5):979-981.
- Gregg AR, Skotko BG, Benkendorf JL, et al. Noninvasive prenatal screening for fetal aneuploidy, 2016 update: a position statement of the American College of Medical Genetics and Genomics. Genet Med. 2016:18(10):1056-1065.
- Wapner RJ, Martin CL, Levy B, et al. Chromosomal microarray versus karyotyping for prenatal diagnosis. N Engl J Med. 2012;367(23):2175-2184.
- Martin KA, Iyengar S, Kalyan A, et al. Clinical experience with a single-nucleotide polymorphism-based noninvasive prenatal test for five clinically significant microdeletions. Clin Genet. 2017 Jul 11. doi: 10.1111/cge.13098. [Epub ahead of print]
Kimberly Martin, MD, FRCSC, FCCMG, FACOG, FACMG, serves as the senior global medical director of Natera’s Women’s Health franchise, and previously served as the medical director for Reproductive Testing at Natera, Inc..
Trudy McKanna, MS, CGS, serves as Medical Science Liaison Manager for Natera, Inc.